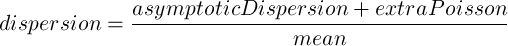
|
(11.1) |
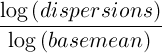
|
(11.2) |
The points used in this model are weight using the normalized mean count.
The DESeq2 method should be used when you have multiple replicates for each of your conditions. At least two replicates are required. Geneious Prime compares expression levels using the DESeq2 package within R (see the R project website), which is automatically downloaded and installed the first time the DESeq method is run on Windows and MacOS. On Linux systems, you will need to install R manually. Refer to 11.2.8
The Fit Type defines the model that will be used by DESeq2 to explain the observed dispersion of read counts:
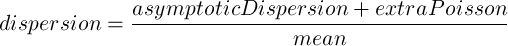
|
(11.1) |
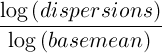
|
(11.2) |
The points used in this model are weight using the normalized mean count.
For more information refer to the estimateDispersions package documentation.
Samples can either be assigned to conditions manually, by selecting either A or B in the Assign Conditions table for each expression track (sample), or automatically based on a field on the expression tracks.
You can select any properties on Expression Level tracks with exactly two values for Geneious to use for automatically assigning conditions. To do this, you first need to set up a property for Geneious to use. This can be done by adding a Meta-data field to the sample reads before assembly or to the sample contigs prior to running Calculate Expression Levels (Geneious will then propagate the information to create a property on the Expression Level track), or by adding a property directly to the Expression Level tracks.
To add a Meta-data field to your samples, go to the Info tab and Create a new Meta-data type in Edit Meta-Data Types, then add a field of this type with Add Meta-Data. For example, you could add a Meta-data field of type ‘Cancer’ with values ‘Yes’ and ‘No’. Meta-data fields must have exactly two values to be used for automatically assigning conditions.
To add a property directly to the expression level tracks, click on the arrow to the left of the track name, choose Edit Fields and then Add a new property.